Mutation Query
| | | | Allele 1: | W312R | | Allele 2: | R574W | Allelic information known | | Refine query |
|
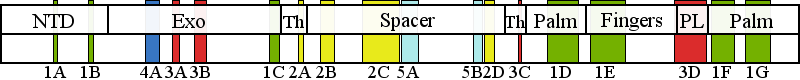
| | | Residue W312 | | Cluster assignment: | | | Cluster description: | Partitioning loop | | Subcluster: | 3B (residues 303-319) | | Subcluster description: | A helix-coil-helix module (residues 295-312) located in the Exo domain that has been termed the "orienter" module | | POLG domain: | Exonuclease domain |
| Residue R574 | | Cluster assignment: | | | Cluster description: | Upstream DNA binding channel | | Subcluster: | 2C (residues 561-617) | | Subcluster description: | Subcluster 2C contains motif 1, which in Pol I was shown to fold into a loop that binds DNA in the channel (Loh and Loeb, 2005). Motif 1, together with subcluster 2D, form the major face of the putative DNA binding channel. | | POLG domain: | Spacer domain |
|
|
Mutation Information
|
| W312R | | | | Number of patients: (with W312R) | 4 | | Found as the only mutation: | 25% of entries (1 patient) | | Non-allelic with: | W312R (50%) R574W (25%) |  | Show Patient Data |
| Patient data are sorted by mutation combination frequency. | Reference: | Di Fonzo et al, 2003; | | Description: | PEO, sensory-motor polyneuropathy, dysphagia. | | Mutations: | W312R, W312R | | Age group: | adult | | Age of Onset: 57, Age of Patient: n/a, Age of Death: n/a |
| Reference: | Del Bo et al, 2003; | | Description: | PEO, complicated by dysphagia, myopathy, sensorimotor polyneuropathy and dysphagia. | | Mutations: | W312R, W312R | | Age group: | adult | | Age of Onset: n/a, Age of Patient: 57, Age of Death: n/a |
| Reference: | Horvath et al, 2006; | | Description: | PEO +dysphagia/myopathy. | | Mutations: | R574W, W312R | | Age group: | adult | | Age of Onset: 37, Age of Patient: n/a, Age of Death: n/a |
| Reference: | Agostino et al, 2003; | | Description: | PEO, bilateral ptosis, severe limitation of ocular motility, and a mosaic distribution of ragged-red and cytochrome c oxidase–negative fibers in the muscle biopsy. All patients had multiple deletions of muscle mtDNA. | | Mutations: | W312R | | Age group: | adult | | Age of Onset: 39, Age of Patient: n/a, Age of Death: n/a |
Back to top |
| R574W | | | | Number of patients: (with R574W) | 4 | | Non-allelic with: | A467T (75%) W312R (25%) |  | Show Patient Data |
| Patient data are sorted by mutation combination frequency. | Reference: | Kollberg et al, 2006; | | Description: | The girl was healthy and developed normally until one year of age when, during a mild respiratory infection, she had repeated multifocal partial seizures with clonic jerking and unconsciousness. The episode was later followed by the development of muscular hypotonia and of myoclonus. Mild psychomotor regression, ataxia, and slightly elevated serum transaminases were noticed. At 3 years of age, a mild infection provoked a prolonged status epilepticus, which lasted 3 weeks and included stroke-like features and rightsided hemiparesis. This was followed by a progressive deterioration of cognitive and motor functions and the development of ptosis and external ophthalmoplegia. Alpers. Febrile infections provoked repetitive status epilepticus with seizures in the form of migrating epilepsia partialis continua. mtDNA Deletions. | | Mutations: | A467T, R574W | | Age group: | infantile | | Age of Onset: 1, Age of Patient: 1.5, Age of Death: 4.333 |
| Reference: | Kollberg et al, 2006; | | Description: | mtDNA Deletions. She was healthy until 7 months of age when she, in association with pyelonephritis, had onset of myoclonic seizures and weakness of the left arm and leg. leftsided myoclonic seizures progressing to status epilepticus in the form of migrating EPC and unconsciousness. pneumonia and colitis. | | Mutations: | A467T, R574W | | Age group: | infantile | | Age of Onset: 0.583, Age of Patient: 1, Age of Death: 1.166 |
| Reference: | Spinazzola et al, 2009; | | Description: | Alpers. Alive at 10 years. Sister died at age 27. | | Mutations: | A467T, R574W | | Age group: | childhood | | Age of Onset: 3, Age of Patient: 10, Age of Death: n/a |
| Reference: | Horvath et al, 2006; | | Description: | PEO +dysphagia/myopathy. | | Mutations: | R574W, W312R | | Age group: | adult | | Age of Onset: 37, Age of Patient: n/a, Age of Death: n/a |
Back to top |
|
|
|
|
The following information is based on existing patient data and pathogenic cluster assignment.
Pathogenicity information for a patient with mutations in Clusters 2 and 3: Age of onset information is extracted from a total of 33 patients and/ or patient families. | Age of onset | | |
33- 17- | 14
| 2
| 3
| 14
| | | infantile | childhd | juvenile | adult | | | 42% | 6% | 9% | 42% | |
All mutations mapping within the pathogenic clusters are at high risk for pathogenicity. In general, a patient must have a pathogenic mutation in both of his/ her POLG genes to develop a POLG-related syndrome.  | Symptoms described in patients with cluster3-cluster2 mutations | |
| Symptoms in patients with combination
cluster2:cluster3 | | PEO | 45.5% | | Epilepsy | 27.3% | | Ptosis | 24.2% | | Encephalopathy | 24.2% | | Dysphagia | 21.2% | | Developmental delay | 18.2% | | Myopathy | 15.2% | | Movement disorder (ataxia) | 12.1% | | Ragged red fibers | 12.1% | | Liver failure | 12.1% | | Liver dysfunction | 12.1% | | Lactic acidosis | 9.1% | | Muscle weakness | 9.1% | | Stroke | 9.1% | | Hypotonic | 9.1% | | Dysarthria | 9.1% | | Areflexia | 9.1% | | Status epilepticus | 6.1% | | Myoclonic seizures | 6.1% | | Sensory ataxia | 6.1% | | Ophthalmoplegia | 6.1% | | Failure to thrive | 6.1% | | Psychomotor delay | 6.1% | | GI dysmotility | 6.1% | | +21 other symptoms in under 5.0% of the patients |
| | Data gathered from clinical descriptions for 33 patients |
| Symptoms by group | | CPEO | 57.6% | | CNS symptoms | 36.4% | | Developmental Delay | 36.4% | | Myopathy | 36.4% | | Seizures | 33.3% | | Hepatopathy | 24.2% | | Ataxia | 18.2% | | Other | 18.2% | | Alpers syndrome | 12.1% | | GI symptoms | 12.1% | | Hypotonia | 9.1% | | Neuropathy | 9.1% |
| | [Show grouping information] |
|
|
|